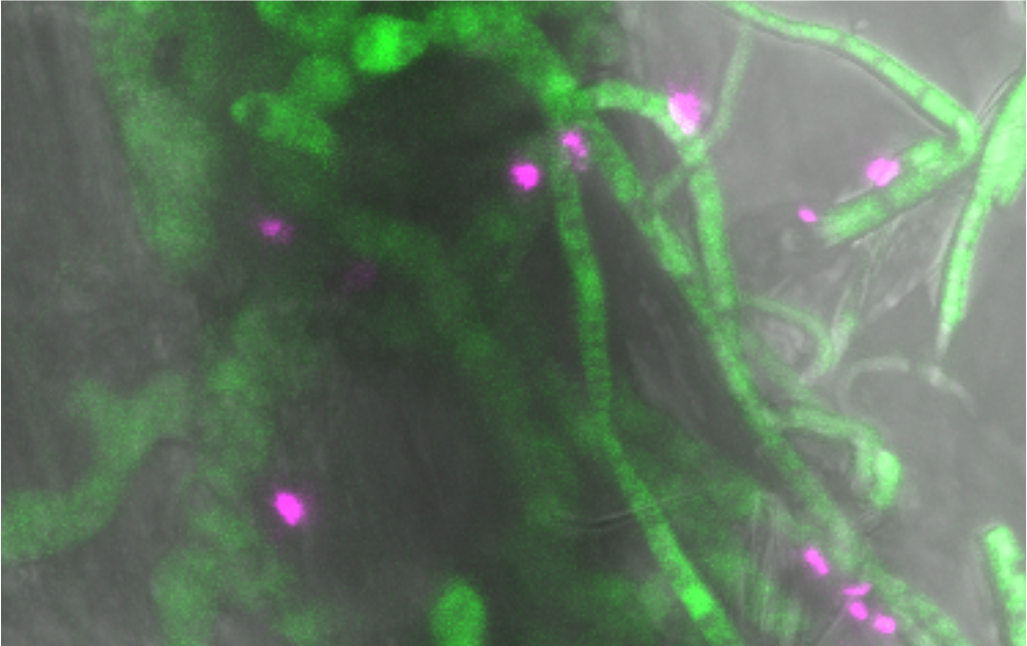
Asymptomatic plants in nature are colonized by a rich diversity of microbes comprising bacteria, fungi, but also protists and viruses (i.e. the plant microbiota). These microorganisms form multi-kingdom microbial consortia that impact plant growth and productivity. Due to strong environmental noise, it remains challenging to understand the organizational rules guiding the establishment of complex (i.e. multi-kingdom) microbial consortia on plant roots as well as their impact on host health in nature. Because diverse microbes are found together in soil and on plant roots, microbiota members likely engage in widespread microbe-microbe interactions, ranging from cooperative to competitive in nature. Therefore, determining the full potential of microbiota effects on plant growth, health and reproduction depends not only on an understanding of the well-studied host-microbe interactions but also on the less characterized microbe-microbe interactions. It is now technically feasible to address these important concepts by combining a) high-throughput microbial profiling data (or metagenomics), b) large-scale microbial isolation and establishment of reference culture collections, and c) microbiota reconstitution experiments with germ-free plants, in order to identify causal relationships between tested variables. The different projects carried out in my group are supported by the Max Planck Society, the European Research Council (ERC starting grant MICRORULES) and the DFG-funded DECRyPT priority programme (Deconstruction and Reconstruction of the Plant Microbiota: SPP 2125).
Research questions
(1) What are the environmental factors that drive the establishment of multikingdom microbial consortia in plant roots?
(2) How do microbe-microbe interactions promote plant health?
(3) What microbial and host genes are important for microbiota establishment in plant roots?
(4) How do plants coordinate microbiota establishment, defense responses, and abiotic stress tolerance?
Current projects in the group
Biogeographical footprint of the Arabidopsis root microbiota on a continental scale (Paloma Duran, Thorsten Thiergart). In collaboration with Carlos Alonso Blanco (Spanish National Biotechnology Centre, Madrid, Spain), Fabrice Roux (French National Institute for Agricultural Research, Toulouse, France), Eric Kemen (Tübingen University, Tübingen, Germany), and Jon Ågren (Uppsala University, Uppsala, Sweden), we investigate structural convergence and divergence of the root microbiota (bacteria, fungi, and oomycetes) on a continental scale. We have selected 17 natural Arabidopsis populations from across Europe and are working to disentangle the contribution of host compartments (soil, rhizosphere, rhizoplane, root), host biogeography, and season to microbial community structure.
Establishment of Arabidopsis root-derived microbial culture collections (Brigitte Pickel, Wolfgang Schmalenbach). The development of sequence-indexed bacterial, fungal, and oomycetal culture collections is a prerequisite for assembling synthetic microbial consortia and represents a key resource for testing the impact of the microbiota on plant fitness. Notably, our culture collections contain a significant proportion of the most abundant root-associated microbes detected by a culture-independent approach have a representative strain in (Bai et al. 2015 Nature, Duran et al. 2018 BioRxiv). Using these collections, we address important ecological principles in microbiota reconstitution experiments.
Impact of microbial inter-kingdom interactions on community assembly and plant health (Paloma Duran). Using gnotobiotic plant systems, we are deciphering the organizational rules that govern the establishment of complex microbial consortia on plant roots and the consequences of inter-kingdom microbe-microbe interactions for plant health. By inoculating microbial groups in these gnotobiotic systems (bacteria, fungi, oomycetes) either separately or in combination, we aim to understand the relevance of cross-kingdom microbe-microbe interactions for microbial community structure and plant health.
Characterization of bacterial genetic determinants conferring plant protection against pathogenic fungi (Felix Getzke). Because bacteria and fungi co-occur in soil and in plant roots, members of these two kingdoms likely engage in widespread microbe-microbe competition. The project’s aims are to establish a high-throughput gnotobiotic plant system to systematically test the effect of bacterial-fungal interactions on plant health and to identify bacterial genes conferring robust pathogen protection. This project will shed new light into how microbial competition maintains fungal-bacterial balance in and outside plant roots.
Whole-genome sequencing and metatranscriptome profiling of complex microbial SynComs (Nathan Vannier, Thorsten Thiergart). We have sequenced the genomes of phylogenetically unrelated microbes (bacteria, fungi and oomycetes) that were successfully isolated from healthy Arabidopsis roots in collaboration with the Max Planck Genome Center and the Joint Genome Institute (1KFG: collaboration with Francis Martin, INRA Nancy, France). By combining large-scale microbial cultivation methods with whole-genome sequencing data, we aim to decipher the genetic determinants governing microbial adaptation to plant roots. These genomic resources will also allow high-resolution metatranscriptome profiling of complex microbial SynComs during root colonization on germ-free plants or upon perturbation.
Role of the plant immune system in controlled accommodation of beneficial microbes (Katarzyna Wolinska). Recent studies suggest that the plant innate immune system must be viewed as a microbial management system also required for interactions with beneficial microbes (Hacquard et al. 2016 Annual review of Phytopathology). We dissect which branches of the plant immune system are required for root colonization by different microbial groups, leading to accurate maintenance of microbiota balance in roots. We are particularly interested to test whether an intact immune system is needed for microbiota-mediated beneficial outcome on Arabidopsis growth in gnotobiotic plant systems.
Definition of functional links coordinating microbiota-dependent tradeoffs between growth and defense under stress conditions (Shiji Hou). The role of the root microbiota on plant growth and its impact on plant immune responses under abiotic stress conditions remains unclear. Particularly, light is one of the fundamental environmental factors underlying plant growth and development, and can also influence plant immune responses. Using a multifaceted approach (synthetic communities, microbial community profiling, Arabidopsis mutants, RNAseq experiments, pathogen assays…) we aim at understanding the interplay between light perception, microbiota establishment, and defense activation in Arabidopsis. This project will facilitate efforts to define and deploy useful microbes that enhance plant performance under suboptimal light conditions.